Default models
Examples of common models are shown below.
[1]:
from comod import models
[2]:
from IPython.display import display, Math
[3]:
def show_graph(model):
return model.plot_tikz(
layout={s: (i, 0) for i, s in enumerate(model.states + [model.nihil_state])},
edge_curved=0.3
)
[4]:
def show_latex(model):
display(Math(model.to_latex()))
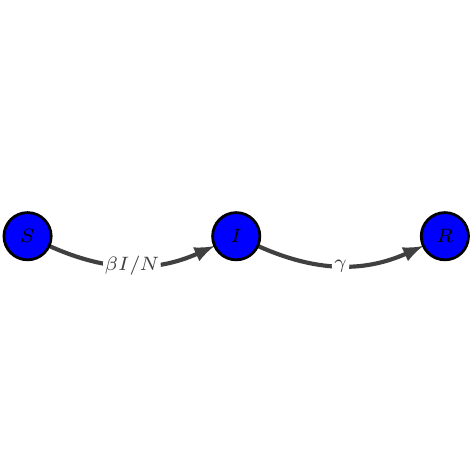
SIR
[5]:
show_graph(models.sir)
[5]:
[6]:
show_latex(models.sir)
$\displaystyle \begin{array}{lcl} \dot{S} &=& -\beta I / N S\\
\dot{I} &=& \beta I / N S - \gamma I\\
\dot{R} &=& \gamma I
\end{array}$
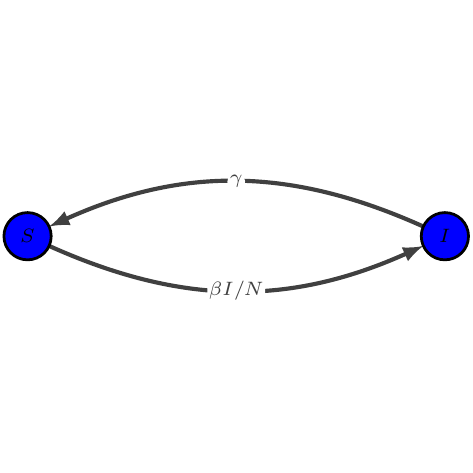
SIS
[7]:
show_graph(models.sis)
[7]:
[8]:
show_latex(models.sis)
$\displaystyle \begin{array}{lcl} \dot{S} &=& -\beta I / N S + \gamma I\\
\dot{I} &=& \beta I / N S - \gamma I
\end{array}$
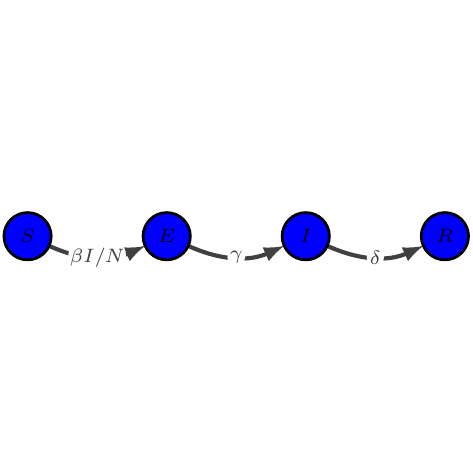
SEIR
[9]:
show_graph(models.seir)
[9]:
[10]:
show_latex(models.seir)
$\displaystyle \begin{array}{lcl} \dot{S} &=& -\beta I / N S\\
\dot{E} &=& \beta I / N S - \gamma E\\
\dot{I} &=& \gamma E - \delta I\\
\dot{R} &=& \delta I
\end{array}$
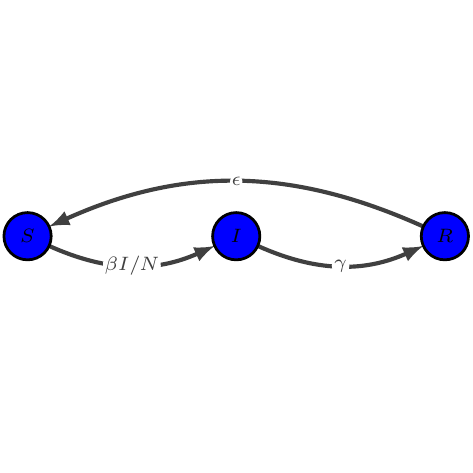
SIRS
[11]:
show_graph(models.sirs)
[11]:
[12]:
show_latex(models.sirs)
$\displaystyle \begin{array}{lcl} \dot{S} &=& -\beta I / N S + \epsilon R\\
\dot{I} &=& \beta I / N S - \gamma I\\
\dot{R} &=& \gamma I - \epsilon R
\end{array}$
Models with vital dynamics
Models with vital dynamics are easilly created extending an existing model.
[13]:
from comod import add_natural
[14]:
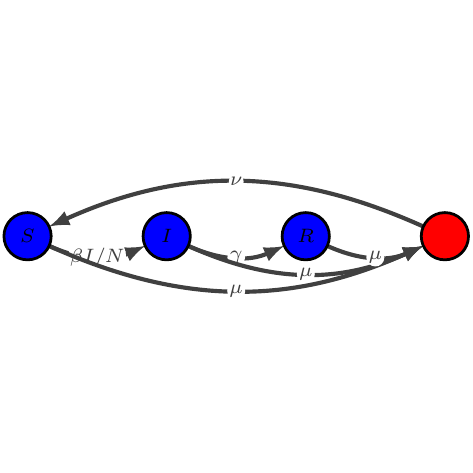
show_graph(add_natural(models.sir))
[14]:
[15]:
show_latex(add_natural(models.sir))
$\displaystyle \begin{array}{lcl} \dot{S} &=& -\beta I / N S + \nu N - \mu S\\
\dot{I} &=& \beta I / N S - \gamma I - \mu I\\
\dot{R} &=& \gamma I - \mu R
\end{array}$
[ ]: